We are thrilled to announce the release of polars-bio, a revolutionary Python DataFrame library designed specifically for large-scale genomics data analysis. After extensive development and optimization, polars-bio is now available on PyPI and ready to transform how genomics researchers work with massive interval datasets.
Genomics research increasingly demands computationally intensive analyses of relationships between genomic features—millions of intervals representing genes, mutations, regulatory elements, and other biological markers across chromosomes. While Python’s Pandas library has long been the workhorse for tabular data manipulation, its memory inefficiency and scalability limitations have become critical bottlenecks for modern genomics workflows. polars-bio addresses these challenges head-on with a fundamentally different architecture built on Rust, Apache DataFusion, and Apache Arrow.
What is polars-bio?
polars-bio is a high-performance Python library that enables fast, parallel, and memory-efficient operations on genomic interval datasets. It combines the power of Polars—a next-generation DataFrame library written in Rust—with specialized genomic operations optimized for interval data. The library’s core components are implemented in Rust using Apache DataFusion as the query engine and Apache Arrow for zero-copy data representation, ensuring maximum efficiency both in computation and memory usage.
Whether you’re working with 100,000 intervals or 100 million, polars-bio is built to scale seamlessly. It maintains compatibility with both Polars and Pandas DataFrame formats, making integration into existing pipelines smooth and straightforward.
Key Features That Matter
Blazing Performance: polars-bio delivers exceptional speed for genomic operations. In real-world benchmarks comparing 1.2 million intervals against 100 intervals, our library achieves:
- Overlap queries: 6.5x faster than alternatives
- Nearest neighbor queries: 15.5x faster
- Count overlaps operations: dramatically optimized
- Coverage calculations: unmatched efficiency
Memory Efficiency: Traditional in-memory DataFrame approaches struggle with large genomic datasets. polars-bio’s Apache Arrow backend provides zero-copy data exchange and native out-of-core streaming capabilities, allowing you to process data that exceeds your available RAM.
Genomics-First API: The library includes popular genomic operations as first-class DataFrame methods, eliminating the need for iteration or custom implementations. Perform interval overlaps, nearest neighbor searches, and coverage calculations with simple, intuitive API calls.
SQL-Powered Processing: For complex data manipulations, leverage the native SQL query engine powered by Apache DataFusion. Write complex queries without leaving Python, combining bioinformatic logic with standard SQL operations.
Cloud-Native Architecture: Process large-scale genomics data stored in cloud environments (S3, GCS) without materializing everything into memory. Built on Apache OpenDAL, polars-bio enables federated and streamed data access from cloud storage, essential for modern distributed genomics workflows.
Parallel Execution: Take advantage of multi-core processors with native parallel operations. Our benchmarks demonstrate the dramatic speedups achieved when scaling across multiple threads.
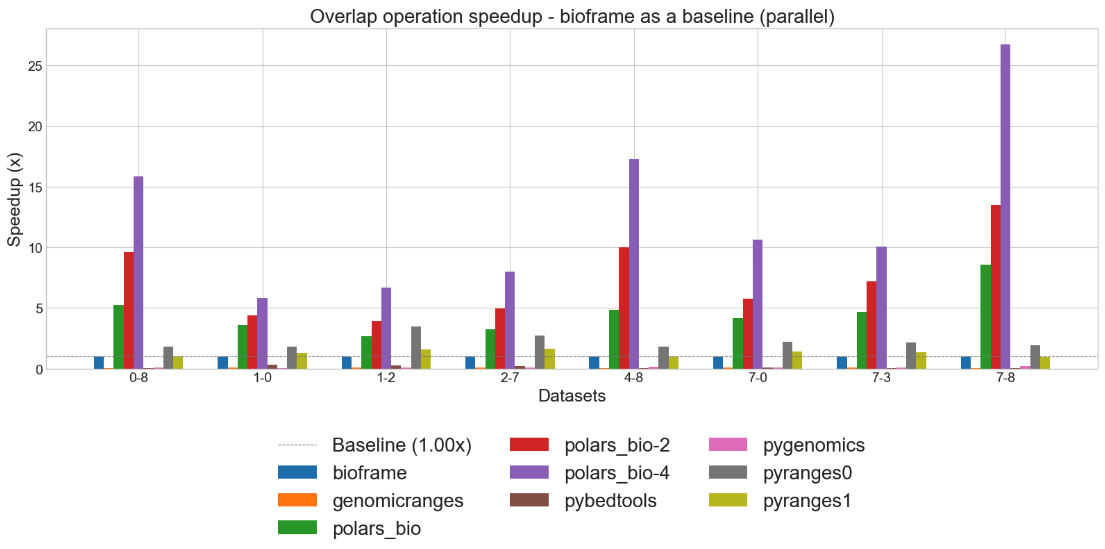
Figure: Parallel overlap query performance demonstrating dramatic scaling efficiency across multiple cores—essential for processing large genomic datasets in production environments.
Comprehensive File Format Support: Work with diverse bioinformatics file formats (BAM, VCF, BED, GFF, and more) thanks to integration with the noodles library. Read genomic data directly into optimized DataFrames without conversion steps.
Cross-Platform Availability: Pre-built wheel packages are available for Linux, Windows, and macOS (both ARM64 and x86_64 architectures), ensuring you can use polars-bio regardless of your development environment.
Why This Matters for Genomics
Genomic analysis is fundamentally about understanding relationships between intervals. Finding overlaps between mutations and regulatory regions, calculating coverage across genomic features, identifying nearest genes—these are operations that seem simple in concept but become computationally intensive at scale. A typical GWAS study might compare millions of variants against thousands of genomic features. With Pandas-based workflows, such analyses could take hours or require distributed computing infrastructure.
polars-bio brings production-grade performance to individual researchers’ laptops and labs’ servers. You can now tackle datasets that previously required Hadoop clusters or specialized HPC infrastructure. This democratization of genomic analysis empowers smaller research groups to compete with well-resourced centers.
The library also shines in exploratory data analysis. Rapid iteration on hypotheses—testing different interval filters, thresholds, and aggregations—becomes feasible when queries return results in milliseconds rather than minutes.
Getting Started
Installation is straightforward via pip:
pip install polars-bioFor detailed installation options and dependency information, visit the quick start guide. We provide comprehensive tutorials, an API reference, and a full feature documentation.
Performance Benchmarks
For a comprehensive overview of performance improvements across different operations and scales, check out our detailed performance benchmarks page. We provide transparent comparisons, methodology details, and guidelines for optimizing your specific use cases.
Citation
If polars-bio contributes to your research, please cite the peer-reviewed publication:
Wiewiorka M, Khamutou P, Zbysiński M, Gambin T. polars-bio—fast, scalable and out-of-core operations on large genomic interval datasets. Bioinformatics, 2025; btaf640. doi:10.1093/bioinformatics/btaf640
Looking Forward
polars-bio represents a significant leap forward for genomics research. By combining Rust performance, Apache’s ecosystem, and community-driven bioinformatics expertise, we’ve created a tool that meets the demands of modern genomic science.
We invite you to explore polars-bio, test it with your datasets, and join the community. Visit the full documentation, check out the GitHub repository, or join our Discord community to connect with other users and contributors.
The future of genomic data analysis is here. Experience it for yourself.
